- Research Article
- Open access
- Published:
Polynomial-Time Algorithm for Controllability Test of a Class of Boolean Biological Networks
EURASIP Journal on Bioinformatics and Systems Biology volume 2010, Article number: 210685 (2010)
Abstract
In recent years, Boolean-network-model-based approaches to dynamical analysis of complex biological networks such as gene regulatory networks have been extensively studied. One of the fundamental problems in control theory of such networks is the problem of determining whether a given substance quantity can be arbitrarily controlled by operating the other substance quantities, which we call the controllability problem. This paper proposes a polynomial-time algorithm for solving this problem. Although the algorithm is based on a sufficient condition for controllability, it is easily computable for a wider class of large-scale biological networks compared with the existing approaches. A key to this success in our approach is to give up computing Boolean operations in a rigorous way and to exploit an adjacency matrix of a directed graph induced by a Boolean network. By applying the proposed approach to a neurotransmitter signaling pathway, it is shown that it is effective.
1. Introduction
Various approaches to modeling, analysis, and control synthesis of biological networks such as gene regulatory networks and metabolic networks have been recently developed in the control community as well as the theoretical biology community [1]. In these approaches, it is one of the final goals to develop systematic drug discovery and cancer treatment [2, 3]. Biological networks in general can be expressed by ordinary/partial differential equations with high nonlinearity and high dimensionality. Since such complexities cause difficulties in analysis and control design, various simpler models such as Petri nets, Bayesian networks, Boolean networks, and hybrid systems have been proposed for dealing with complex and large-scale biological networks at the expense of rigorous analysis (see e.g., [4, 5]).
This paper discusses the controllability problem of biological networks. In gene regulatory networks, for example, the controllability problem is defined as the problem of determining whether expressions of genes of interest can be arbitrarily controlled by expressions of a specified set of the other genes. As far as we know, two approaches to the controllability analysis of such biological networks have been developed so far: a piecewise affine model-based approach and a Boolean network model-based approach. However, the former approach can be applied to only the class of relatively low-dimensional systems [6, 7].
On the other hand, a Boolean network model, where binary state variables are assigned to nodes and the transition rules of the state are given by Boolean functions [8, 9], will be more practical for analysis of large-scale biological networks thanks to its bold simplification. Akutsu et al. have recently discussed the controllability problem of Boolean networks with control nodes and controlled nodes and have proven that this problem is NP-hard in a general setting [10]. Furthermore, they have proposed a polynomial-time algorithm for the classes of networks including a tree structure or at most one loop, and an exponential-time algorithm for the other classes. Indeed there is a criticism that a Boolean network model is too simple as a model of biological networks, but for large-scale networks it will be able to provide some indication or clue towards further detailed analysis. Thus various approaches based on this model have been well-studied so far (see e.g., [11–19]).
Motivated by the theoretical results in [10], this paper also focuses on the controllability problem of Boolean networks with control nodes and controlled nodes and proposes a sufficient condition for the Boolean network to be controllable, which can be easily verified by a polynomial-time algorithm. Our standing point is to give up computing complex Boolean operations in a rigorous way and to focus on deriving an easily-checkable sufficient condition for controllability so as to be applied to large-scale networks. The obtained algorithm is based on simple operations on an adjacency matrix of a directed graph induced by a Boolean network. This is a remarkable point of our approach, different from the method in [10], and enables us to apply our approach to a wider class of Boolean networks including nontree structures.
First, after the definition of controllability of Boolean network models with control nodes and controlled nodes is described, a sufficient condition for the controllability is derived in the form of an algorithm. Next, the computational complexity for the algorithm is discussed to show that it is a polynomial-time algorithm. In addition, PC-based numerical experiments show that the obtained algorithm is applicable to a class of Boolean networks with at least 1000 nodes. Finally, as an illustrative example, the proposed algorithm is applied to the Boolean network model of a neurotransmitter signaling pathway [20], which expresses an interaction pathway between the glutamatergic and dopaminergic receptors. Note that the polynomial-time algorithm proposed in [10] cannot be always applied to this problem. This Boolean network model consists of 16 nodes, and the problem of simultaneously controlling two important nodes among them, that is, concentration of exocytosis and phospholipase C, is discussed based on the proposed algorithm. As a result, we show that for example, they can be simultaneously controlled by keeping substance concentration at the other 4 nodes constant with appropriate values.
Notation 1. Let  denote the set of nonnegative integers and
denote the set of nonnegative integers and  the set of
the set of  matrices consisting of elements
matrices consisting of elements  and
and  . We also denote by
. We also denote by  and
and  the
the  identity matrix and the
identity matrix and the  zero matrix, respectively. For simplicity of notation, we sometimes use the symbol
zero matrix, respectively. For simplicity of notation, we sometimes use the symbol  instead of
instead of  and the symbol
and the symbol  instead of
instead of  . Let
. Let  express the transpose of the matrix
express the transpose of the matrix  .
.
2. Boolean Network Models
This section provides a brief review on a Boolean network model [8, 9]. A Boolean network model consists of a set of nodes and a set of regulation rules for nodes, where each node expresses a gene, a molecule, or an event in the genetic network. The state variable  at node
at node  takes a Boolean value of
takes a Boolean value of  or
or  representing "inactive" or "active" status of the node, respectively. A regulation rule for each node is given in terms of a Boolean function, and each node state changes synchronously.
representing "inactive" or "active" status of the node, respectively. A regulation rule for each node is given in terms of a Boolean function, and each node state changes synchronously.
As an example, we consider a very simple and interesting Boolean network model of an apoptosis network in Figure 1 given by

where  ,
,  , and
, and  denote logical NOT, AND, and OR, respectively,
denote logical NOT, AND, and OR, respectively,  denotes the discrete time, the concentration level (high or low) of the tumor necrosis factor (TNF, a stimulus) is denoted by
denotes the discrete time, the concentration level (high or low) of the tumor necrosis factor (TNF, a stimulus) is denoted by  , the concentration level of the inhibitor of apoptosis proteins (IAP) by
, the concentration level of the inhibitor of apoptosis proteins (IAP) by  , the concentration level of the active caspase 3 (C3a) by
, the concentration level of the active caspase 3 (C3a) by  , and the concentration level of the active caspase 8 (C8a) by
, and the concentration level of the active caspase 8 (C8a) by  . Here if the binary variable
. Here if the binary variable  has the value of "1", then the concentration of a certain reactant gets larger than a prescribed threshold (i.e., it is active), otherwise less than that. In addition, logical NOT corresponds to inhibition of gene expressions.
has the value of "1", then the concentration of a certain reactant gets larger than a prescribed threshold (i.e., it is active), otherwise less than that. In addition, logical NOT corresponds to inhibition of gene expressions.
Since the caspase C3a is responsible for cleaving or breaking many other proteins, a high-level of the C3a concentration, that is,  implies cell near-death; otherwise, cell survival. As seen in (1), if the concentration of IAP is high (
implies cell near-death; otherwise, cell survival. As seen in (1), if the concentration of IAP is high ( ) or the concentration of the caspase C8a is low (
) or the concentration of the caspase C8a is low ( ), then the concentration of C3a gets low, that is,
), then the concentration of C3a gets low, that is,  . On the other hand,
. On the other hand,  and
and  at the next time depend on the value of
at the next time depend on the value of  as well as
as well as  . In this way, some dynamical interactions exist. See [21, 22] for further details.
. In this way, some dynamical interactions exist. See [21, 22] for further details.
A general form of a Boolean network model is given by the state equation

where  is the state vector at time
is the state vector at time  , and
, and  is a Boolean function, where logical operators consist of AND (
is a Boolean function, where logical operators consist of AND ( ), OR (
), OR ( ), NOT (
), NOT ( ), and XOR (
), and XOR ( ).
).
3. Problem Formulation
In a Boolean network model (2), the state  is uniquely determined by giving the initial state
is uniquely determined by giving the initial state  , which implies that (2) is an autonomous system and has no control nodes.
, which implies that (2) is an autonomous system and has no control nodes.
On the other hand, this paper will consider the Boolean network model with control (i.e., input) nodes and controlled (i.e., output) nodes to discuss the output-controllability of this model. This model is given by

where each element of  denotes the state of the control node whose value can be arbitrarily given as an external control input in the Boolean network, each element of
denotes the state of the control node whose value can be arbitrarily given as an external control input in the Boolean network, each element of  denotes the state of the node except for the control nodes in the Boolean network, and each element of
denotes the state of the node except for the control nodes in the Boolean network, and each element of  denotes the state of the node to be controlled as an output in the network. Note here that
denotes the state of the node to be controlled as an output in the network. Note here that  does not imply a measured output. Hereafter according to control theory,
does not imply a measured output. Hereafter according to control theory,  ,
,  , and
, and  are called a "state", "control input" and "output", respectively. In addition,
are called a "state", "control input" and "output", respectively. In addition,  is a Boolean function, and
is a Boolean function, and  is the output matrix satisfying for each element
is the output matrix satisfying for each element  of
of 

Furthermore, the product of  and
and  in
in  expresses a product operation on matrices/vectors of the real number field. Thus the above condition on
expresses a product operation on matrices/vectors of the real number field. Thus the above condition on  guarantees that the output is the state variable itself, that is, for each
guarantees that the output is the state variable itself, that is, for each  there exists
there exists  such that
such that  holds. The case of
holds. The case of  is also included here. This condition on
is also included here. This condition on  will not be restrictive in analyzing controllability of biological networks such as gene regulatory networks, since the relation on regulation among genes/molecules will be mainly discussed there.
will not be restrictive in analyzing controllability of biological networks such as gene regulatory networks, since the relation on regulation among genes/molecules will be mainly discussed there.
For the system  of (3), the notion of output-controllability is defined as follows.
of (3), the notion of output-controllability is defined as follows.
Definition 1. Suppose that for the system  of (3), the finite time
of (3), the finite time  and the initial state
and the initial state  are given. Then the system
are given. Then the system  is said to be
is said to be  -output-controllable at
-output-controllable at  if for every
if for every  , there exists a control input sequence
, there exists a control input sequence  ,
,  , such that
, such that  . Furthermore, the system
. Furthermore, the system  is said to be
is said to be  -output-controllable if it is
-output-controllable if it is  -output-controllable at every
-output-controllable at every  .
.
The above notion of controllability comes from the fact that, for example, in control of genetic networks we often would like to determine if expressions of certain gene of interest (corresponding to  ) will be able to be inhibited (or activated) by means of appropriately adjusting the expressions of a given set of genes (corresponding to
) will be able to be inhibited (or activated) by means of appropriately adjusting the expressions of a given set of genes (corresponding to  ). It is remarked that we assume that the control time
). It is remarked that we assume that the control time  is explicitly specified in the above definition.
is explicitly specified in the above definition.
Let us get back to the Boolean network model (1) of an apoptosis network. As discussed in [21, 22], we consider  (TNF) itself as a control input. So by ignoring the dynamics on
(TNF) itself as a control input. So by ignoring the dynamics on  , that is,
, that is,  , we suppose in (1) that
, we suppose in (1) that  and
and  , which yields (3) of the form
, which yields (3) of the form

where  denotes the
denotes the  -th element of
-th element of  . As for the output
. As for the output  , either case of
, either case of

can be treated by assumption. Then let us verify the  -output controllability of the system (5). As discussed in Section 2,
-output controllability of the system (5). As discussed in Section 2,  expresses cell near-death, and
expresses cell near-death, and  expresses cell survival. So we would like to know if the system is
expresses cell survival. So we would like to know if the system is  -output-controllable with respect to the output
-output-controllable with respect to the output  . Suppose that
. Suppose that  (i.e., the initial states of IAP, C3a and C8a are all low-level),
(i.e., the initial states of IAP, C3a and C8a are all low-level),  (i.e.,
(i.e.,  ), and
), and  . Then since
. Then since  holds independently of
holds independently of  by simple calculation, we see that system (5) is not
by simple calculation, we see that system (5) is not  -output-controllable at
-output-controllable at  , which implies that we cannot control the system from the state "cell survival" within 2 time steps no matter how the control value of
, which implies that we cannot control the system from the state "cell survival" within 2 time steps no matter how the control value of  is given.
is given.
On the other hand, suppose in (1) that  and
and  . Then we obtain (3) of the form
. Then we obtain (3) of the form

where  is ignored. Suppose that
is ignored. Suppose that  ,
,  , and
, and

(i.e.,  ). Then since
). Then since  and
and  are obtained, we see that system (5) is
are obtained, we see that system (5) is  -output-controllable at
-output-controllable at  , for example, (a)
, for example, (a)  for
for  ,
,  , (b)
, (b)  for
for  ,
,  , (c)
, (c)  for
for  ,
,  , and (d)
, and (d)  for
for  ,
,  . This implies we can simultaneously control the value of
. This implies we can simultaneously control the value of  and
and  at
at  . In this way, the proposed controllability enables us to verify the existence of a control input sequence such that the output has the desired value in a given finite time, and the obtained result indicates how to give the value of a control input sequence.
. In this way, the proposed controllability enables us to verify the existence of a control input sequence such that the output has the desired value in a given finite time, and the obtained result indicates how to give the value of a control input sequence.
Next, we will explain our basic strategy for deriving the controllability condition. Let us consider a Boolean network expressed as the state equation

which is given by [10]. Although this model is very simple, it provides significant clues to address this problem. For the Boolean network model (9), we can consider three possible specifications, choosing either  ,
,  , or
, or  to be the control input for the system.
to be the control input for the system.
First, suppose that  and
and  , that is,
, that is,  itself is the control input. Then it follows that
itself is the control input. Then it follows that

Note here that  is ignored because we assume that
is ignored because we assume that  itself is the control input. As for the output
itself is the control input. As for the output  , either case of
, either case of  ,
,  ,
,  can be considered in this case. Consider the controllability of the system (10) with
can be considered in this case. Consider the controllability of the system (10) with  (i.e.,
(i.e.,  ) for
) for  . In this example, we will consider whether system (10) is
. In this example, we will consider whether system (10) is  -output-controllable or not by directly calculating state trajectories of each system. From (10), we have
-output-controllable or not by directly calculating state trajectories of each system. From (10), we have

So if  ,
,  holds irrespective of the value of
holds irrespective of the value of  , similarly for the case of
, similarly for the case of  . Therefore, we see that system (10) is not
. Therefore, we see that system (10) is not  -output-controllable. In the same way, we see that system (10) is not
-output-controllable. In the same way, we see that system (10) is not  -output-controllable in every case of
-output-controllable in every case of  ,
,  , and
, and  for
for  .
.
Secondly, suppose that  and
and  , that is,
, that is,  itself is regarded as the control input. Then we obtain
itself is regarded as the control input. Then we obtain

where  is ignored. Consider the controllability of the system (12) for
is ignored. Consider the controllability of the system (12) for  . From (12) we have
. From (12) we have

Thus we see that the system is not  -output-controllable for
-output-controllable for  , while that the system is
, while that the system is  -output-controllable with
-output-controllable with  for both cases of
for both cases of  and
and  .
.
Finally, suppose that  and
and  . Then we obtain
. Then we obtain

where  is ignored. Consider the system (14) with
is ignored. Consider the system (14) with  . From (14), we have
. From (14), we have

which implies that system (14) is  -output-controllable. However, in the case of
-output-controllable. However, in the case of  , we have
, we have

which implies that the system (14) is not  -output-controllable.
-output-controllable.
Note that for (11) with  , we see that the controllability property does not hold due to the fact that
, we see that the controllability property does not hold due to the fact that  directly depends on
directly depends on  . On the other hand, for (13) with
. On the other hand, for (13) with  ,
,  is adjacent to
is adjacent to  and
and  in the Boolean network, which implies that
in the Boolean network, which implies that  is arbitrarily given by
is arbitrarily given by  and
and  . In a similar way,
. In a similar way,  is adjacent to
is adjacent to  . However,
. However,  cannot be realized by
cannot be realized by  and
and  because
because  always holds when
always holds when  . These examples are very important in discussing the controllability in a Boolean network, that is, if the Boolean function of
. These examples are very important in discussing the controllability in a Boolean network, that is, if the Boolean function of  includes an initial state
includes an initial state  , or includes the same input in the outputs at the same time, then the system in question is not
, or includes the same input in the outputs at the same time, then the system in question is not  -output-controllable. In the following section, by motivating the above discussion, we will consider to derive a controllability condition.
-output-controllable. In the following section, by motivating the above discussion, we will consider to derive a controllability condition.
Remark 1. In the above example, we assume that when some genes are identified as control inputs, the original dynamics of the corresponding genes can be ignored. However, in the case that the corresponding gene has a strong interaction with other genes, this assumption may not be suitable. One of methods for coping with such a case is to add a new gene (node) that works as the control input [10], where it is called an external control node. Our approach below can be also applied to this case.
4. Output-Controllability Condition
4.1. Preliminaries
This section presents a sufficient condition for the system (3) to be  -output-controllable in the form of an algorithm.
-output-controllable in the form of an algorithm.
Consider a simple example given by

This system has the following relation:

Similarly, we see that  ,
,  , hold identically. In Boolean functions, identical equations are in general given by
, hold identically. In Boolean functions, identical equations are in general given by

where  is any Boolean function of a vector of binary variables. Obviously such identities on
is any Boolean function of a vector of binary variables. Obviously such identities on  or
or  affect the controllability in a Boolean network (note that even if
affect the controllability in a Boolean network (note that even if  ,
,  holds irrespective of
holds irrespective of  ).
).
Let us consider again the Boolean network model (5) of an apoptosis network. If we suppose that  ,
,  (i.e.,
(i.e.,  ), and
), and  , then by a simple calculation, we obtain the following identity:
, then by a simple calculation, we obtain the following identity:

So in Boolean biological networks, there exists the case that identities are appeared. However, identities may not be appeared in the real biological relevance. The reasons why such identities are appeared are that the state is binarized and that a time-delay of the state is ignored. To overcome the latter point, a temporal Boolean network model  has been proposed in [23]. However, identities may appear even in a temporal Boolean network. The output-controllability condition proposed below can be similarly applied to a temporal Boolean network model.
has been proposed in [23]. However, identities may appear even in a temporal Boolean network. The output-controllability condition proposed below can be similarly applied to a temporal Boolean network model.
Thus first of all, we will focus on finding such identities in  before discussing a kind of initial condition and a kind of input-independency. This will require the introduction for several symbols.
before discussing a kind of initial condition and a kind of input-independency. This will require the introduction for several symbols.
The following assumption is made.
Assumption 1. The Boolean function  in (3) has no redundant variables.
in (3) has no redundant variables.
For example, in the logical function  ,
,  holds. So
holds. So  is a redundant variable, and
is a redundant variable, and  can be rewritten as
can be rewritten as  . Any given Boolean function can be changed so as to satisfy Assumption 1: after it is transformed into an appropriate canonical form (e.g., Reed-Muller canonical form (polynomials over the finite field
. Any given Boolean function can be changed so as to satisfy Assumption 1: after it is transformed into an appropriate canonical form (e.g., Reed-Muller canonical form (polynomials over the finite field  )), it is easy to eliminate redundant variables by expanding based on four operations over
)), it is easy to eliminate redundant variables by expanding based on four operations over  . Also in the identification of Boolean network models (e.g., see [24]), since the correlations between variables are checked, the Boolean function
. Also in the identification of Boolean network models (e.g., see [24]), since the correlations between variables are checked, the Boolean function  in (3) will satisfy Assumption 1 in many cases. By Assumption 1, it is guaranteed that the Boolean function
in (3) will satisfy Assumption 1 in many cases. By Assumption 1, it is guaranteed that the Boolean function  itself does not include any identities, although
itself does not include any identities, although  may include some identities. Let
may include some identities. Let  denote the number of the logical NOT appeared in (3), where the logical NOT operators are distinguished when the corresponding terms are different even if the corresponding variables are the same. In addition, consider the fictitious inputs
denote the number of the logical NOT appeared in (3), where the logical NOT operators are distinguished when the corresponding terms are different even if the corresponding variables are the same. In addition, consider the fictitious inputs  ,
,  , which have one-to-one correspondence with the variables operated by the logical NOT, that is,
, which have one-to-one correspondence with the variables operated by the logical NOT, that is,  or
or  in (3). Then the system (3) can be equivalently rewritten as the following system:
in (3). Then the system (3) can be equivalently rewritten as the following system:

where the Boolean function  does not include the logical NOT, and
does not include the logical NOT, and

For example, system (17) is rewritten as

subject to  .
.
Next, consider the adjacency matrix  for the directed graph induced by the Boolean network of the system (21). For example, the adjacency matrix for the system (23) is given by
for the directed graph induced by the Boolean network of the system (21). For example, the adjacency matrix for the system (23) is given by

where if there exists an arc from node  to node
to node  , then the
, then the  -th element of
-th element of  is
is  . Hereafter, without loss of generality, the
. Hereafter, without loss of generality, the  -th element of
-th element of  is assigned to node
is assigned to node  in the directed graph, where
in the directed graph, where  . In the case of (24),
. In the case of (24),  ,
,  ,
,  ,
,  , and
, and  are assigned to nodes
are assigned to nodes  ,
,  ,
,  ,
,  , and
, and  , respectively. Then in Figure 2, which shows a temporal/spatial network of the system (17), we say that for example, there exists a path between
, respectively. Then in Figure 2, which shows a temporal/spatial network of the system (17), we say that for example, there exists a path between  and
and  .
.
Using the adjacency matrix  , we also compute the matrix
, we also compute the matrix  ,
,  , where
, where

In the case of the system (17), we have

where  .
.
For the system (21),  expresses whether there exist paths between
expresses whether there exist paths between  and
and  ,
,  and
and  , or
, or  and
and  for any given
for any given  . In the case of (26), we see that
. In the case of (26), we see that  is adjacent to
is adjacent to  and
and  . In other words,
. In other words,  expresses which elements of
expresses which elements of  ,
,  , and
, and  are variables of a Boolean function representing
are variables of a Boolean function representing  . However, note here that from
. However, note here that from  , we cannot specify an explicit form of the Boolean function in question.
, we cannot specify an explicit form of the Boolean function in question.
Furthermore, the following symbol is used:

where  ,
,  , and
, and  . Let also
. Let also  ,
,  , and
, and  denote each element of
denote each element of  ,
,  ,
,  , respectively. If
, respectively. If  holds, then there exist
holds, then there exist  paths between
paths between  and
and  . For the state
. For the state  ,
,  , of the system (3), let
, of the system (3), let  express the index set of elements of
express the index set of elements of  operated by the logical NOT as
operated by the logical NOT as  . In a similar way, for the control input
. In a similar way, for the control input  ,
,  , of the system (3), let
, of the system (3), let  express the index set of elements of
express the index set of elements of  operated by the logical NOT as
operated by the logical NOT as  . Here,
. Here,  holds. In addition, there is a one-to-one correspondence between each element of
holds. In addition, there is a one-to-one correspondence between each element of  ,
,  and the index
and the index  of
of  . Let
. Let  and
and  express the index
express the index  of
of  corresponding to
corresponding to  and
and  , respectively. In the case of the system (23),
, respectively. In the case of the system (23),  ,
,  hold, and for
hold, and for  ,
,  holds.
holds.
Finally, we define the following matrices:

where

4.2. Proposed Algorithm
Now we are in a position to propose a  -output-controllability test algorithm. Since this kind of problem is NP-hard [10], we pay our attention on deriving a sufficient condition for the controllability. Although this sufficient condition is given in the form of an algorithm, it is somewhat complex. Thus before describing an algorithm, we describe the outline of the algorithm.
-output-controllability test algorithm. Since this kind of problem is NP-hard [10], we pay our attention on deriving a sufficient condition for the controllability. Although this sufficient condition is given in the form of an algorithm, it is somewhat complex. Thus before describing an algorithm, we describe the outline of the algorithm.
First, we consider a necessary condition for  to include identical equations. From Figure 2 of the example (23), we see that
to include identical equations. From Figure 2 of the example (23), we see that  in (18), which has no identities, has two paths from
in (18), which has no identities, has two paths from  , and that
, and that  is connected to some node on the paths. In this way, if some identical equation exists in
is connected to some node on the paths. In this way, if some identical equation exists in  , there always exist more than 2 paths from
, there always exist more than 2 paths from  to some state and also the logical-NOT operations exist on the paths, which is a necessary condition and not necessarily a sufficient condition. Since it will spend huge time to rigorously specify the existence of identities for a large network, we consider here to exclude the cases satisfying the above necessary condition, that is, we do not determine here the controllability in such cases.
to some state and also the logical-NOT operations exist on the paths, which is a necessary condition and not necessarily a sufficient condition. Since it will spend huge time to rigorously specify the existence of identities for a large network, we consider here to exclude the cases satisfying the above necessary condition, that is, we do not determine here the controllability in such cases.
Next, for the system that includes no identical equations, we use a kind of input-independency to determine the controllability. For example, consider the case that neither identity on  nor
nor  exists in
exists in  and that
and that  is expressed by
is expressed by

as a result of recursive calculation (see Section 6 for such an example), where  ,
,  are some Boolean functions. This system is obviously
are some Boolean functions. This system is obviously  -output controllable because each
-output controllable because each  is expressed by different
is expressed by different  and no
and no  exists in
exists in  . From the viewpoint of adjacency relation, this implies that there exists no path between
. From the viewpoint of adjacency relation, this implies that there exists no path between  and
and  , there exists at least one path from each
, there exists at least one path from each  to some
to some  , and each
, and each  has a path with only one
has a path with only one  or has no path to any
or has no path to any  . This can be easily found from the adjacency matrix, although it is a sufficient condition for the controllability. This is a rough story of our approach.
. This can be easily found from the adjacency matrix, although it is a sufficient condition for the controllability. This is a rough story of our approach.
The proposed algorithm is given as follows.
Algorithm 1 (T-output-controllability test algorithm).
Part A: Check of the Existence of Identical Equations.
Step 1. Set  . Compute
. Compute  ,
,  , and
, and  .
.
Step 2. If  , go to Step 6. Otherwise set
, go to Step 6. Otherwise set  . Compute
. Compute  ,
,  , and
, and  .
.
Step 3. If there exists  such that
such that  or
or  such that
such that  , denote them by
, denote them by  or
or  , respectively, and go to Step 4. Otherwise, go to Step 2 if
, respectively, and go to Step 4. Otherwise, go to Step 2 if  and go to Step 6 if
and go to Step 6 if  .
.
Step 4. If there exists  such that
such that  or
or  such that
such that  , and
, and  1 or
1 or  1 holds, go to Step 8. Otherwise, go to Step 5.
1 holds, go to Step 8. Otherwise, go to Step 5.
Step 5.
Substep 5.1. Set  .
.
Substep 5.2. If any element of  -th column or
-th column or  -th column in
-th column in  is greater than or equal to
is greater than or equal to  , go to Step 8. Otherwise, go to Substep 5.3.
, go to Step 8. Otherwise, go to Substep 5.3.
Substep 5.3. If  , set
, set  and go to Substep 5.2, or else go to Step 2.
and go to Substep 5.2, or else go to Step 2.
Part B: Check of the Independence of Each
 .
.
Step 6. If the following conditions hold for the matrices  and
and  in (28), system (3) is
in (28), system (3) is  -output-controllable, or else if only condition (i) does not hold, then go to Step 7. Otherwise go to Step 8.
-output-controllable, or else if only condition (i) does not hold, then go to Step 7. Otherwise go to Step 8.
-
(i)
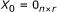 holds;
holds; -
(ii)
each column vector of
 is a nonzero vector;
is a nonzero vector; -
(iii)
each row vector of
 is a zero vector, or has only one element with a nonzero value.
is a zero vector, or has only one element with a nonzero value.
Step 7. Suppose  for a given constant vector
for a given constant vector  . Let
. Let  denote the index set of elements of
denote the index set of elements of  that are constant for any
that are constant for any  (
( ). Then if the following condition holds, system (3) is
). Then if the following condition holds, system (3) is  -output-controllable at
-output-controllable at  . Otherwise, go to Step 8.
. Otherwise, go to Step 8.
-
(iv)
For
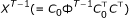 , there exists no
, there exists no 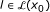 satisfying
satisfying 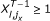 .
.
Step 8. This algorithm cannot determine whether the system (3) is  -output-controllable or not (at
-output-controllable or not (at  ).
).
The above algorithm allows us to determine the  -output-controllability of the system as follows.
-output-controllability of the system as follows.
First, noting that the identical equations have the form in (19), and  is obtained recursively from (21), we see that the identical equations appeared in
is obtained recursively from (21), we see that the identical equations appeared in  always have the form
always have the form


where  denotes either variable of
denotes either variable of  or
or  ,
,

and  ,
,  are some subsets of the index set
are some subsets of the index set  . Then using the forms of (31) and (32), the following lemma on Part A of Algorithm 1 is obtained.
. Then using the forms of (31) and (32), the following lemma on Part A of Algorithm 1 is obtained.
Lemma 1.
In Step 6,
 includes neither identities of (31) nor identities of (32).
includes neither identities of (31) nor identities of (32).
Proof. In Step 3, from  for some
for some  ,
,  , and
, and  , we see that more than 2 paths from
, we see that more than 2 paths from  to
to  exist, which is necessary for the identity on
exist, which is necessary for the identity on  to exist (similarly for the case of
to exist (similarly for the case of  ). Thus we next focus on the existence of logical NOT (i.e.,
). Thus we next focus on the existence of logical NOT (i.e.,  ) in these paths in Step 4 and Step 5.
) in these paths in Step 4 and Step 5.
Consider the case that the logical NOT (i.e.,  ) corresponding to
) corresponding to  or
or  obtained in Step 3 exists in (21), in other words, either
obtained in Step 3 exists in (21), in other words, either  or
or  holds. Then the condition
holds. Then the condition  implies that the term
implies that the term  is included in the paths in question, which is a necessary condition for the existence of the identity in
is included in the paths in question, which is a necessary condition for the existence of the identity in  . Thus we exclude this case (Step 4). (similarly for the case
. Thus we exclude this case (Step 4). (similarly for the case  ).
).
In the other case, from (31), (32), for  , some
, some  , to exist in the paths in question is necessary for the existence of identities. If any element of the
, to exist in the paths in question is necessary for the existence of identities. If any element of the  -column or the
-column or the  -column of
-column of  is greater than or equal to 1, some element of
is greater than or equal to 1, some element of  exists in the paths in question. Thus we exclude this case (Substep 5.2). Therefore, it follows that
exists in the paths in question. Thus we exclude this case (Substep 5.2). Therefore, it follows that  includes no identities in Step 6.
includes no identities in Step 6.
From Lemma 1, we see that the case that  includes the identities that have the form of (31) or (32) is excluded from the viewpoint of a necessary condition for the identity to exist in
includes the identities that have the form of (31) or (32) is excluded from the viewpoint of a necessary condition for the identity to exist in  . Thus we obtain the following theorem.
. Thus we obtain the following theorem.
Theorem 1.
For a given
 , the following statements hold.
, the following statements hold.
-
(i)
the system (3) is
 -output-controllable if conditions (i), (ii), and (iii) in Step 6 hold subject to Part A of Algorithm 1,
-output-controllable if conditions (i), (ii), and (iii) in Step 6 hold subject to Part A of Algorithm 1,
-
(ii)
for a given
 , the system (3) is
, the system (3) is
 -output-controllable at
-output-controllable at
 if condition (iv) in Step 7 holds subject to Part A and Step 6.
if condition (iv) in Step 7 holds subject to Part A and Step 6.
Proof. First, the statement (i) is proven for the system satisfying the condition that  includes neither identities of (31) nor identities of (32). From Lemma 1, this condition is satisfied in Step 6. Then condition (i) in Step 6 implies that there exists no path between each element of
includes neither identities of (31) nor identities of (32). From Lemma 1, this condition is satisfied in Step 6. Then condition (i) in Step 6 implies that there exists no path between each element of  and each element of
and each element of  , since the
, since the  -th element of
-th element of  expresses if a path from
expresses if a path from  to
to  exists or not. On the other hand, note that
exists or not. On the other hand, note that  -th element of
-th element of  expresses if a path from
expresses if a path from  to
to  exists or not
exists or not  . Thus condition (ii) in Step 6 implies that there exists at least one path from each element of
. Thus condition (ii) in Step 6 implies that there exists at least one path from each element of  to some
to some  .
.
Furthermore, condition (iii) in Step 6 means that the input  for each
for each  and
and  has a path connected to only one element of
has a path connected to only one element of  or has no path to any element of
or has no path to any element of  . From these conditions, it follows that each
. From these conditions, it follows that each  affects at most one
affects at most one  and not the other
and not the other  ,
,  . Hence the value of
. Hence the value of  can be independently specified by the corresponding
can be independently specified by the corresponding  , which implies that system (21) is
, which implies that system (21) is  -output-controllable.
-output-controllable.
Next, the statement (ii) is proven. Since condition (i) in Step 6 does not hold, in this case, there exists a path between some element of  and some element of
and some element of  . On the other hand, condition (iv) in Step 7 guarantees that there exists no path between constant elements of
. On the other hand, condition (iv) in Step 7 guarantees that there exists no path between constant elements of  and elements of
and elements of  . Thus
. Thus  is not affected by the value of
is not affected by the value of  . Therefore, from (ii)–(iv), it follows that system (21) is
. Therefore, from (ii)–(iv), it follows that system (21) is  -output-controllable at
-output-controllable at  . This completes the proof.
. This completes the proof.
As an example, consider system (17) again. Suppose  . The matrices
. The matrices  ,
,  ,
,  of Step 1 are given by (26), and
of Step 1 are given by (26), and  ,
,  ,
,  of Step 2 are
of Step 2 are

In Step 3, from  , we obtain
, we obtain  and
and  . In Step 4, from
. In Step 4, from  and
and  , we have
, we have  and
and  . So go to Step 8, that is, it is impossible to determine if system (17) is
. So go to Step 8, that is, it is impossible to determine if system (17) is  -output-controllable. In fact, from (18),
-output-controllable. In fact, from (18),  includes the identity
includes the identity  . Thus we see that there exists an identical equation.
. Thus we see that there exists an identical equation.
Let us also consider the case of  ,
,  in the system (17). Then for
in the system (17). Then for  , we have
, we have

From Step 1 Step 2
Step 2 Step 3
Step 3 Step 6, we can see that the system (17) is
Step 6, we can see that the system (17) is  -output-controllable. In fact, by simple calculation, the Boolean function of
-output-controllable. In fact, by simple calculation, the Boolean function of  is derived as
is derived as  .
.
As for identical equations, the proposed algorithm excludes the case of  as well as (19). This is a weak point of this algorithm. Furthermore, consider the following system:
as well as (19). This is a weak point of this algorithm. Furthermore, consider the following system:

This system is  -output-controllable for
-output-controllable for  . However, the proposed algorithm cannot determine whether this system is
. However, the proposed algorithm cannot determine whether this system is  -output-controllable or not; thus there exists a class of systems such that the proposed algorithm cannot determine the controllability. Needless to say, it will not be so easy to cope with various cases stated above due to high nonlinearity of Boolean functions.
-output-controllable or not; thus there exists a class of systems such that the proposed algorithm cannot determine the controllability. Needless to say, it will not be so easy to cope with various cases stated above due to high nonlinearity of Boolean functions.
While the proposed algorithm includes such disadvantages, one of the main advantages of the algorithm is that the computational complexity of the above algorithm is very small. This will be discussed in the following section.
5. Computational Complexity Analysis
In this section, we discuss the computational complexity of the algorithm proposed in the previous section.
First, let us recall the definition of the symbols used here. The number of the state, the control input, and the output in (3) are denoted by  ,
,  , and
, and  , respectively. The number of the logical NOT appeared in (3) is expressed by
, respectively. The number of the logical NOT appeared in (3) is expressed by  . In addition,
. In addition,  expresses the control time. Then the following result is obtained.
expresses the control time. Then the following result is obtained.
Lemma 2.
The computational complexity of the proposed algorithm is
 for
for
 ,
,
 .
.
Proof. The computation of the proposed algorithm consists of (a) checking each condition of Part A, and (b) checking whether conditions (i) to (iv) hold or not.
First, (b) is considered. The computational complexity to compute  is
is  . So the computational complexity to compute
. So the computational complexity to compute  and
and  is given by both
is given by both  . Further, the computational complexity to compute the product of
. Further, the computational complexity to compute the product of  and
and  is
is  . So by simple calculation, the computational complexity of
. So by simple calculation, the computational complexity of  is obtained as
is obtained as  . The computational complexity of generating
. The computational complexity of generating  is obviously less than the case of
is obviously less than the case of  . Therefore, the computational complexity to compute
. Therefore, the computational complexity to compute  and
and  is
is  , which also includes the computational complexity to check conditions (i) to (iv) in Steps 6 and 7 for given
, which also includes the computational complexity to check conditions (i) to (iv) in Steps 6 and 7 for given  and
and  .
.
Next, (a) is considered. The matrices  ,
,  ,
,  are obtained directly from
are obtained directly from  , and the computational complexity of Step 5 is
, and the computational complexity of Step 5 is  . As a result, since the computational complexity of each checking in Part A is
. As a result, since the computational complexity of each checking in Part A is  , the computational complexity of Part A is less than
, the computational complexity of Part A is less than  .
.
Therefore, the computational complexity of the proposed algorithm is given by  .
.
From Lemma 2, we see that the proposed algorithm is a polynomial-time algorithm. Furthermore, the computational time for performing the proposed algorithm is evaluated by numerical experiments, where the total computational time in Part B is measured because from the proof of Lemma 2 we see that the computational complexity of Part B is dominant. So the adjacency matrices to be evaluated are generated randomly for each  , where
, where  ,
,  are given. The results are shown in Table 1, where MATLAB on the computer with the Intel Core 2 Duo CPU 3.0 GHz and the 2 GB memory is used. In Table 1, the worst computational time implies the worst value among
are given. The results are shown in Table 1, where MATLAB on the computer with the Intel Core 2 Duo CPU 3.0 GHz and the 2 GB memory is used. In Table 1, the worst computational time implies the worst value among  cases randomly selected for each
cases randomly selected for each  . From Table 1, we see that the proposed algorithm can be applied to relatively large-scale Boolean network models.
. From Table 1, we see that the proposed algorithm can be applied to relatively large-scale Boolean network models.
 )
)6. Application to Neurotransmitter Signaling Pathway
In this section, the proposed algorithm is applied to a Boolean network model of interaction pathway between the glutamatergic and dopaminergic receptors in Figure 3, which has been proposed in [20]. In this pathway, exocytosis, by which a cell directs the contents of secretory vesicles out of the cell membrane, is regulated, depending on the value of neurotransmitters such as dopamine and glutamate. Then it is important from the viewpoint of synaptic plasticity to consider whether exocytosis can be controlled by regulating other elements. In the Boolean network model of Figure 3, the dopamine (neurotransmitter,  ) is synthesized by tyrosine hydroxylase (
) is synthesized by tyrosine hydroxylase ( ) and catabolized by COMT (
) and catabolized by COMT ( ). The dopamine binds to the dopamine receptor 1 (DRD1,
). The dopamine binds to the dopamine receptor 1 (DRD1,  ) and the dopamine receptor 2 (DRD2,
) and the dopamine receptor 2 (DRD2,  ). DRD1 stimulates adenylate cyclase
). DRD1 stimulates adenylate cyclase  to activate protein kinase A (
to activate protein kinase A ( ), which activates DARPP32 (
), which activates DARPP32 ( ). DARPP32 inhibits protein phosphatase 1 (
). DARPP32 inhibits protein phosphatase 1 ( ). By inhibitation of protein phosphatase 1, activation of protein kinase A, and presence of the glutamate (
). By inhibitation of protein phosphatase 1, activation of protein kinase A, and presence of the glutamate ( ), the glutamate receptor
), the glutamate receptor  is activated to elevate the concentration of the intracellular calcium (
is activated to elevate the concentration of the intracellular calcium ( ). On the other hand, DRD2 inactivates adenylate cyclase and activates phospholipase C (
). On the other hand, DRD2 inactivates adenylate cyclase and activates phospholipase C ( ) in order to elevate the concentration of the intracellular calcium. The intracellular calcium activates calcineurin (
) in order to elevate the concentration of the intracellular calcium. The intracellular calcium activates calcineurin ( ), which inhibits DARPP32. Also, the intracellular calcium activates packaging proteins (
), which inhibits DARPP32. Also, the intracellular calcium activates packaging proteins ( ) and finally exocytosis (
) and finally exocytosis ( ). The process of exocytosis of the glutamate receptor expresses one of events in synaptic plasticity, that is, if exocytosis is activated, then the neurotransmitter is secreted out of the cell membrane. In this model, the concentration of the above reactants is expressed by a binary variable
). The process of exocytosis of the glutamate receptor expresses one of events in synaptic plasticity, that is, if exocytosis is activated, then the neurotransmitter is secreted out of the cell membrane. In this model, the concentration of the above reactants is expressed by a binary variable  , that is,
, that is,  if it is high, otherwise
if it is high, otherwise  . Then the state equations of this system are given as
. Then the state equations of this system are given as

From Figure 3, we see that this Boolean network includes at least four loops, for example, the loop of  ,
,  ,
,  , and
, and  , the loop of
, the loop of  ,
,  ,
,  ,
,  , and
, and  , and so forth. In synaptic plasticity, it is required that the binary value of
, and so forth. In synaptic plasticity, it is required that the binary value of  expressing exocytosis can be arbitrarily controlled. Furthermore, phospholipase C (
expressing exocytosis can be arbitrarily controlled. Furthermore, phospholipase C ( ) is a kind of enzymes that cleaves phospholipids and as a result protein kinase C as well as calcium
) is a kind of enzymes that cleaves phospholipids and as a result protein kinase C as well as calcium  are activated. The former, protein kinase C, which works outside of the network in Figure 3, is one of key enzymes in signal transduction pathways. Thus since phospholipase C affects the other significant network, it will be important to simultaneously control the value of
are activated. The former, protein kinase C, which works outside of the network in Figure 3, is one of key enzymes in signal transduction pathways. Thus since phospholipase C affects the other significant network, it will be important to simultaneously control the value of  and the value of
and the value of  . Therefore
. Therefore  and
and  are regarded as the output, that is,
are regarded as the output, that is,  . In addition, we assume that
. In addition, we assume that  and
and  cannot be directly controlled.
cannot be directly controlled.
For a fixed dimension of  and the fixed output
and the fixed output  , all combinations of
, all combinations of  ,
,  , are considered as the control inputs, which we call the input-combinations. Then for a given
, are considered as the control inputs, which we call the input-combinations. Then for a given  , the proposed algorithm is applied to the system of the form (3) obtained for each input-combination of
, the proposed algorithm is applied to the system of the form (3) obtained for each input-combination of  . It is remarked that depending on the choice of the kind of control inputs, there exist several cases to which the polynomial-time algorithm proposed in [10] cannot be applied due to the graph-structure constraints. Furthermore, it is also remarked that even for fixed control inputs, the controllability problem is NP-hard. So the problem of finding efficient control inputs that make the system controllable is further harder than this problem.
. It is remarked that depending on the choice of the kind of control inputs, there exist several cases to which the polynomial-time algorithm proposed in [10] cannot be applied due to the graph-structure constraints. Furthermore, it is also remarked that even for fixed control inputs, the controllability problem is NP-hard. So the problem of finding efficient control inputs that make the system controllable is further harder than this problem.
By applying our algorithm to the case of each input-combination of  and each fixed
and each fixed  , we obtain, for example, the following results. In the case of
, we obtain, for example, the following results. In the case of  and
and  , we can find that among
, we can find that among  input-combinations, there exist at least 6 input-combinations of
input-combinations, there exist at least 6 input-combinations of  that make the system
that make the system  -output-controllable. In this way, since the proposed algorithm for each input-combination is very efficient, for example, the computation time via the proposed algorithm is about 10 [sec] for Boolean networks with 600 nodes (see Table 1) and
-output-controllable. In this way, since the proposed algorithm for each input-combination is very efficient, for example, the computation time via the proposed algorithm is about 10 [sec] for Boolean networks with 600 nodes (see Table 1) and  , it enables us to verify the controllability condition for a certain number of input-combinations within a practical time; for example, about 3 [hours] will be required for 1000 input-combinations of a Boolean network with 600 nodes.
, it enables us to verify the controllability condition for a certain number of input-combinations within a practical time; for example, about 3 [hours] will be required for 1000 input-combinations of a Boolean network with 600 nodes.
In the case of  and
and  , we can also find controllable control inputs among
, we can also find controllable control inputs among  input-combinations. For example, we obtain as one of combinations of
input-combinations. For example, we obtain as one of combinations of  and
and  that make the system
that make the system  -output-controllable
-output-controllable


It is remarked that the polynomial-time algorithm proposed in [10] cannot be applied to the system with the state (38) and the input (39) because the network includes the two loops, that is, the loop of  ,
,  ,
,  , and
, and  , and the loop of
, and the loop of  ,
,  ,
,  , and
, and  . Furthermore, based on the above result the Boolean function of
. Furthermore, based on the above result the Boolean function of  can be derived as
can be derived as


which implies that the value of  can be freely given by control inputs, for example, (a)
can be freely given by control inputs, for example, (a)  for
for  ,
,  ,
,  ,
,  ,
,  , and (b)
, and (b)  for
for  ,
,  ,
,  ,
,  ,
,  .
.
Finally, we discuss the control input sequence realizing the desired output values. One of criticisms in control of Boolean networks is to assume that the value of the control input can be arbitrarily given at each time. In many biological systems, this assumption is not always satisfied, and input constraints are frequently imposed. One of input constraints is that the value of the control input is given as a constant within a certain sufficiently long time period. Although it is one of future works to explicitly deal with such an input constraint, based on the proposed algorithm, we may also find a constant-valued sequence of control inputs for which the desired values of outputs are obtained. For example, in (40) and (41), let us consider to find a control input sequence satisfying  and
and  . Since
. Since  ,
,  ,
,  ,
,  , and
, and  are obtained as one of solutions, it is remarked that the following control inputs are given as any binary value:
are obtained as one of solutions, it is remarked that the following control inputs are given as any binary value:  ,
,  ,
,  ,
,  , and
, and  ,
,  ,
,  ,
,  ,
,  , and
, and  ,
,  ,
,  ,
,  ,
,  , and
, and  ,
,  ,
,  ,
,  ,
,  . This allows us to give the values of the control input sequences as a constant, that is,
. This allows us to give the values of the control input sequences as a constant, that is,  ,
,  ,
,  ,
,  ,
,  . Thus the proposed algorithm helps us to find a practically useful control input sequence. Furthermore, this kind of degree of freedom in control inputs may be used for the optimal control problem. Once we can determine control input variables by our algorithm, we can use a tool for finding optimal control input sequences, which have been developed in hybrid control theory, for example, [25, 26]. It is expected that such an analysis will provide one of guidelines in experimental approaches to the control problem of biological networks.
. Thus the proposed algorithm helps us to find a practically useful control input sequence. Furthermore, this kind of degree of freedom in control inputs may be used for the optimal control problem. Once we can determine control input variables by our algorithm, we can use a tool for finding optimal control input sequences, which have been developed in hybrid control theory, for example, [25, 26]. It is expected that such an analysis will provide one of guidelines in experimental approaches to the control problem of biological networks.
7. Conclusion
In this paper, the controllability analysis for biological networks expressed by a Boolean network model with control nodes (inputs) and controlled nodes (outputs) has been discussed. First, a sufficient condition for the Boolean network model to be output-controllable has been derived by exploiting an adjacency matrix of its network graph. The obtained condition, which is given in the form of an algorithm, can be checked in polynomial time with respect to the state/input dimensions and the control time period; it will be one of the powerful tools that can provide some clues for finding effective control inputs to control a large-scale biological network. Next, by PC-based numerical experiments, it has been shown that the proposed method is applicable to large-scale Boolean networks with at least 1000 nodes. Finally, the proposed method has been applied to the Boolean network model expressing a neurotransmitter signaling pathway, and has shown that it is controllable with respect to both exocytosis and phospholipase C when appropriate control inputs are used.
There are many interesting open problems to be addressed in the future. It is one of the most important issues to characterize a class of Boolean networks to which our algorithm can be applied as a necessary and sufficient condition. In addition, extensions to the case of systems with input constraints and uncertainty are also one of the significant topics.
References
Khammash M, Tomlin CJ, Vidyasagar M (Eds): Joint special issue on systems biology In IEEE Transactions on Automatic Control & IEEE Transactions on Circuits and Systems I 2008.
Kitano H: Computational systems biology. Nature 2002, 420(6912):206-210. 10.1038/nature01254
Kitano H: Cancer as a robust system: implications for anticancer therapy. Nature Reviews Cancer 2004, 4(3):227-235. 10.1038/nrc1300
Ferrari-Trecate G, Lygeros J: Workshop on Hybrid Systems Biology. Proceedings of the 45th IEEE Conference on Decision and Control/Workshop Hybrid Systems Biology, 2006
De Jong H: Modeling and simulation of genetic regulatory systems: a literature review. Journal of Computational Biology 2002, 9(1):67-103. 10.1089/10665270252833208
Azuma S-I, Yanagisawa E, Imura J-I: Controllability analysis of biosystems based on piecewise-affine systems approach. IEEE Transactions on Automatic Control 2008, 53(1):139-152.
Belta C, Schug J, Dang T, Kumar V, Pappas GJ, Rubin H, Dunlap P: Stability and reachability analysis of a hybrid model of luminescence in the marine bacterium Vibrio fischeri . Proceedings of the 40th IEEE Conference on Decision and Control (CDC '01), 2001 869-874.
Kauffman SA: Metabolic stability and epigenesis in randomly constructed genetic nets. Journal of Theoretical Biology 1969, 22(3):437-467. 10.1016/0022-5193(69)90015-0
Kauffman SA: The Origins of Order: Self-Organization and Selection in Evolution. Oxford University Press, Oxford, UK; 1993.
Akutsu T, Hayashida M, Ching W-K, Ng MK: Control of Boolean networks: hardness results and algorithms for tree structured networks. Journal of Theoretical Biology 2007, 244(4):670-679. 10.1016/j.jtbi.2006.09.023
Aldana M: Boolean dynamics of networks with scale-free topology. Physica D 2003, 185(1):45-66. 10.1016/S0167-2789(03)00174-X
Datta A, Choudhary A, Bittner ML, Dougherty ER: External control in Markovian genetic regulatory networks. Machine Learning 2003, 52(1-2):169-191.
Datta A, Choudhary A, Bittner ML, Dougherty ER: External control in Markovian genetic regulatory networks: the imperfect information case. Bioinformatics 2004, 20(6):924-930. 10.1093/bioinformatics/bth008
Fauré A, Naldi A, Chaouiya C, Thieffry D: Dynamical analysis of a generic Boolean model for the control of the mammalian cell cycle. Bioinformatics 2006, 22(14):e124-e131. 10.1093/bioinformatics/btl210
Langmead CJ, Jha SK: Symbolic approaches to finding control strategies in Boolean networks. Proceedings of the 6th Asia-Pacific Bioinformatics Conference, 2008 307-319.
Martin S, Zhang Z, Martino A, Faulon J-L: Boolean dynamics of genetic regulatory networks inferred from microarray time series data. Bioinformatics 2007, 23(7):866-874. 10.1093/bioinformatics/btm021
Pal R, Datta A, Dougherty ER: Optimal infinite-horizon control for probabilistic Boolean networks. IEEE Transactions on Signal Processing 2006, 54(6):2375-2387.
Shmulevich I, Dougherty ER, Kim S, Zhang W: Probabilistic Boolean networks: a rule-based uncertainty model for gene regulatory networks. Bioinformatics 2002, 18(2):261-274. 10.1093/bioinformatics/18.2.261
Shmulevich I, Zhang W: Binary analysis and optimization-based normalization of gene expression data. Bioinformatics 2002, 18(4):555-565. 10.1093/bioinformatics/18.4.555
Gupta S, Bisht SS, Kukreti R, Jain S, Brahmachari SK: Boolean network analysis of a neurotransmitter signaling pathway. Journal of Theoretical Biology 2007, 244(3):463-469. 10.1016/j.jtbi.2006.08.014
Chaves M: Methods for qualitative analysis of genetic networks. Proceedings of the European Control Conference, 2009 671-676.
Tournier L, Chaves M: Uncovering operational interactions in genetic networks using asynchronous Boolean dynamics. Journal of Theoretical Biology 2009, 260(2):196-209. 10.1016/j.jtbi.2009.06.006
Silvescu A, Honavar V: Temporal Boolean network models of genetic networks and their inference from gene expression time series. Complex Systems 2001, 13: 54-71.
Akutsu T, Miyano S, Kuhara S: Identification of genetic networks from a small number of gene expression patterns under the boolean network model. Proceedings of the Pacific Symposium on Biocomputing, 1999 17-28.
Bemporad A, Morari M: Control of systems integrating logic, dynamics, and constraints. Automatica 1999, 35(3):407-427. 10.1016/S0005-1098(98)00178-2
Nakayama H, Tanaka H, Ushio T: The formulation of the control of an expression pattern in a gene network by propositional calculus. Journal of Theoretical Biology 2006, 240(3):443-450. 10.1016/j.jtbi.2005.10.014
Acknowledgment
The authors would like to thank Professor Tatsuya Akutsu, Kyoto University for fruitful discussions and valuable comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Kobayashi, K., Imura, JI. & Hiraishi, K. Polynomial-Time Algorithm for Controllability Test of a Class of Boolean Biological Networks. J Bioinform Sys Biology 2010, 210685 (2010). https://doi.org/10.1155/2010/210685
Received:
Accepted:
Published:
DOI: https://doi.org/10.1155/2010/210685


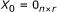 holds;
holds; is a nonzero vector;
is a nonzero vector; is a zero vector, or has only one element with a nonzero value.
is a zero vector, or has only one element with a nonzero value.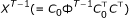 , there exists no
, there exists no 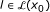 satisfying
satisfying 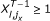 .
. -output-controllable if conditions (i), (ii), and (iii) in Step 6 hold subject to Part A of Algorithm 1,
-output-controllable if conditions (i), (ii), and (iii) in Step 6 hold subject to Part A of Algorithm 1,
 , the system (3) is
, the system (3) is
 -output-controllable at
-output-controllable at
 if condition (iv) in Step 7 holds subject to Part A and Step 6.
if condition (iv) in Step 7 holds subject to Part A and Step 6.
